Main navigation
Faculty research areas
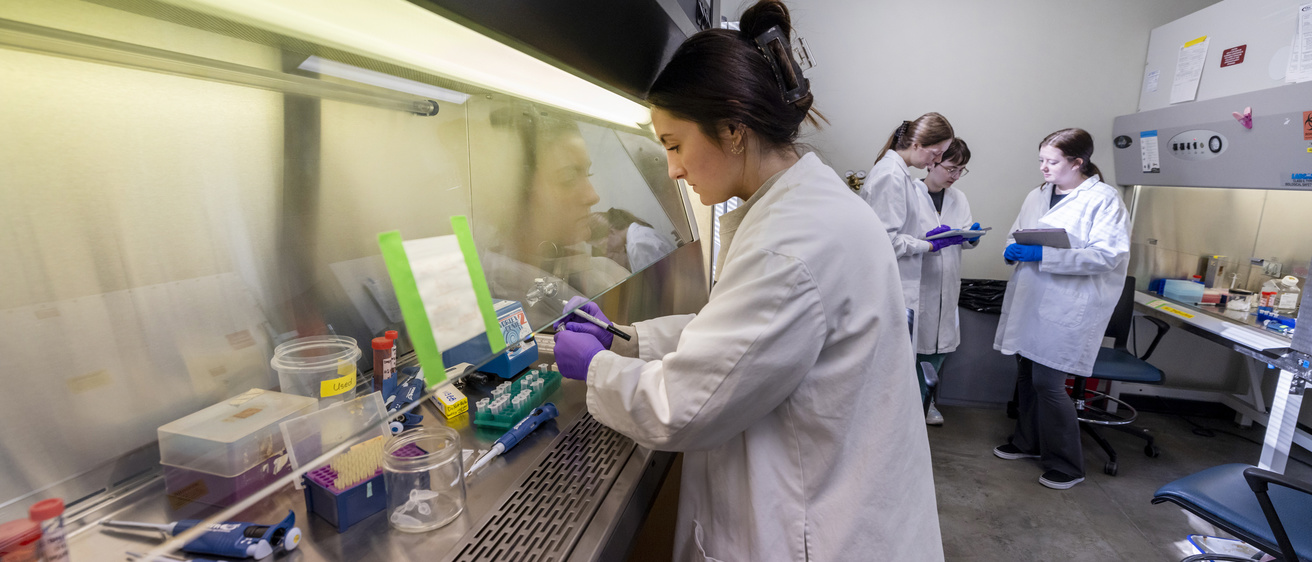
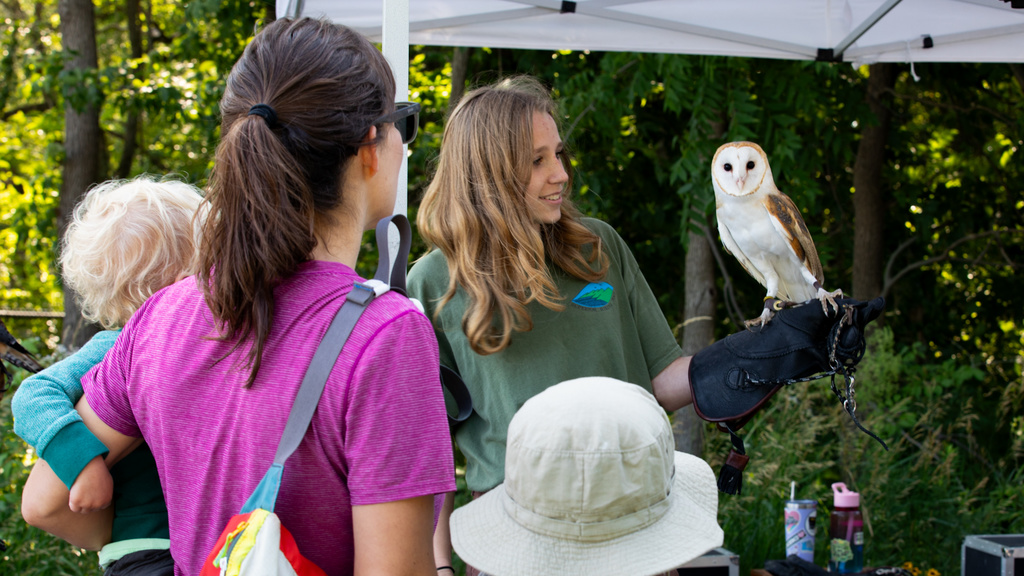
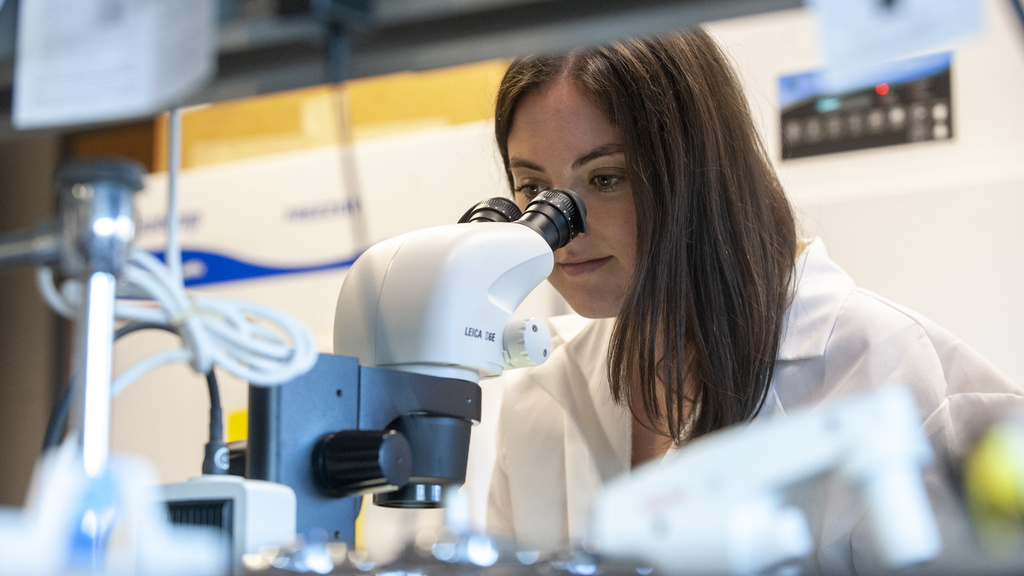
Research in the Department of Biology – both current faculty and future hires – connect to three integrative and cross-disciplinary areas: Modes and Mechanisms of Reproduction; Resilience and Adaptation; and Dynamics of Cells.
Each of these research areas are pursued by faculty across the four disciplines of Biology:

“Iowa allows students to participate in a variety of extracurricular activities. As a first-year, I dipped my toes in many different areas and found my true passion—research. In this field, my mentors were my biggest assets.”
Research news
Biology Researchers trace genetic agent in life-threatening fungal disease
Manak Lab Links Brain's Immune System to Epilepsy
Biology Researchers Discover Genes That Can Cause Cleft Lip and Palate
Biology Professor Finds Genes Associated with Cleft Lip and Palate
Biology Professor Receives $1.3 grant to prevent noise-induced neurodegeneration
Biology professor conducting research on mosquito hearing at Nagoya University in Japan
Interdisciplinary team receives $500K to study climate, environment, and health
Biology professors receive funding to study stress-induced epigenetic changes
Research and creative production in the College of Liberal Arts and Sciences
As part of a top-tier, AAU-accredited public research university, the College of Liberal Arts and Sciences holds scholarly, scientific, and artistic discovery at the heart of our mission.
Throughout our departments and programs, our professors are at the forefront of their disciplines. They bring their world-class research and artistry into their classrooms, studios, and labs, giving students the unparalleled opportunity to learn right from the source of the latest innovations in knowledge and practice.
Graduate students and many undergraduates work side-by-side with faculty members, conducting breakthrough research and creative production that advances humanity’s understanding of ourselves and the complex, ever-changing world in which we live.
The creation of knowledge and understanding is an exhilarating and never-ending mission—and is at the core of every University of Iowa liberal arts and sciences education.